Today is a special day for us at Truveta. In just under two weeks, our team was able to ask and answer an important medical question using one of the largest comprehensive real-time datasets of fully vaccinated Americans.
This inaugural custom study is an illustration of the potential before us. More than the science alone, which we’re also quite excited about, we’re energized by the opportunity that the Truveta Platform can offer to physicians like me and clinical researchers around the world to discover solutions to health problems at new levels of speed and accuracy.
We are fully committed to transparency – sharing the methodology and limitations of the data. As we grow with each new health system that joins Truveta, the dataset in the platform will enable research of rarer conditions and precision treatments, accelerating insights needed for better patient care.
At Truveta, we are working hard to build a platform that will enable researchers to find cures faster, empower every clinician to be an expert, and help families make the most informed decisions about their care. Our vision is saving lives with data, and today represents a first step toward that goal.
Below is our full scientific analysis and approach for our first findings on COVID-19. This reflects early learnings and we have only just begun, but already interesting patterns have emerged. We look forward to sharing more in the future.
-Nick
COVID-19 breakthrough infection and hospitalization in people with comorbidities
From Truveta’s Clinical Researchers
- Nick Stucky, MD, PhD, Director of Clinical and Scientific Research at Truveta and practicing infectious diseases physician and researcher at Providence Portland Medical Center
- Michael Simonov, MD, Director of Clinical Informatics at Truveta, a Health Informatics Lecturer at Yale University School of Medicine and a practicing physician for VA Connecticut.
- Michael Wang, MD, Director of Clinical Informatics at Truveta and assistant professor practicing hospital medicine at University of California San Francisco Health.
- Senthil K. Nachimuthu, MD, PhD, FAMIA, Senior Director of Clinical Informatics at Truveta and an adjunct assistant professor and research associate at University of Utah School of Medicine.
- Peter Smits, PhD, Senior Applied Scientist at Truveta and an expert in Bayesian data analysis with extensive experience in generalized linear models (e.g. regression), hierarchical/multilevel models, survival analysis, longitudinal data analysis, and discrete time series analysis.
- Samuel Gratzl, PhD, Data Visualization Researcher at Truveta and a research software engineer with Carnegie Mellon University.
The questions we asked
Reports of declining vaccine effectiveness against COVID-19 have driven discussion about booster vaccinations (1,2). On September 24, 2021, Pfizer booster vaccinations were approved based on data showing waning vaccine immunity. On October 21, 2021, CDC expanded its recommendation to include boosters for Moderna and Janssen vaccines.
These decisions were based largely on declines in vaccine effectiveness in the elderly population in both U.S. and Israeli populations (3). The CDC recommended booster vaccinations for people age ≥50 with high-risk medical conditions while stating that people age 18-49 with high-risk medical conditions may receive the vaccine based on limited unpublished CDC data (4).
Nevertheless, only limited data were available to understand the risk of breakthrough infections in people with high-risk medical conditions in the U.S. A breakthrough infection is a COVID-19 infection despite being fully vaccinated.
Prior studies in unvaccinated populations have shown more severe outcomes for COVID-19 infection for people with certain high-risk comorbidities such as diabetes, chronic kidney disease (CKD), or who are immunocompromised, e.g., because of cancer or HIV (5–8). Before the SARS-CoV-2 delta variant became prevalent, one study of 108,720 mostly male U.S. veterans with high-risk conditions showed vaccine effectiveness for preventing infection remained high (>95%) but did not evaluate the risk of severe disease (9). Another study in a similar population did not show an elevated risk of severe outcomes in breakthrough infections (10). In order to better understand the risk of breakthrough infection and severe outcomes in high-risk populations, we used the Truveta Platform to ask whether vaccinated patients with comorbidities (CKD, chronic lung disease, diabetes, and those who are immunocompromised) have higher rates of breakthrough COVID-19 infection and higher rates of hospitalization following breakthrough infection than in the general population.
Main findings
Our analysis found that the incidence of COVID-19 breakthrough infection and hospitalization following breakthrough infection was significantly greater among patients with select comorbid medical conditions when compared to the general population. Specifically, people with diabetes, chronic lung disease or CKD have increased incidence of breakthrough infection compared to the general population after adjusting for age, sex, and race. This is not surprising given that these conditions are thought to lead to impaired immune function (11–13). This is consistent with studies in unvaccinated people showing higher risk of infection in these populations as well as a study in mostly male U.S. veterans which showed reduced vaccine effectiveness in patients with a high Charlson comorbidity index (9).
In our study, patients with chronic kidney disease (CKD), chronic lung disease, diabetes or who are immunocompromised were nearly twice as likely to be hospitalized than the general vaccinated population. Overall, these findings add to prior studies showing worse outcomes following COVID-19 infection in people who are immunocompromised, with diabetes, CKD, or with chronic lung disease (5–7) and adds additional support for recommendations of booster vaccines given these groups continue to fare worse than the general vaccinated population.
Diving deeper
Baseline characteristics
Our overall study population consists of 1,707,650 fully vaccinated patients from the Truveta Platform. The Methods section (below) describes in more detail how the population was selected and prepared. We assigned to each patient whether they have a specific comorbidity (CKD, chronic lung disease, diabetes, or those who are immunocompromised) or are comorbidity-free. This led to five populations, one for each comorbidity of interest and one comorbidity-free general population. If a patient has multiple comorbidities, they were assigned to each of the corresponding comorbidities sub populations, see also the limitations discussion below for details. Table 1 summarizes the distributions of raw patient counts within each sub population:
| General Population (Comorbidity-free) | Chronic Lung Disease | Chronic Kidney Disease | Diabetes | Immuno- compromised | ||
| N (fully vaccinated patients) | 1,311,016 (100.0%) | 140,866 (100.0%) | 110,295 (100.0%) | 134,667 (100.0%) | 126,386 (100.0%) | |
| Breakthrough Case | 11,738 (0.9%) | 1,584 (1.1%) | 1,677 (1.5%) | 1,442 (1.1%) | 1,071 (0.8%) | |
| Hospitalized for Breakthrough Case | 876 (0.1%) | 257 (0.2%) | 441 (0.4%) | 286 (0.2%) | 215 (0.2%) | |
| Age Group | ||||||
| <18 | 52,260 (4.0%) | 4,685 (3.3%) | 16 (0.0%) | 342 (0.3%) | 237 (0.2%) | |
| 18-49 | 535,217 (40.8%) | 38,199 (27.1%) | 4,669 (4.2%) | 17,920 (13.3%) | 13,939 (11.0%) | |
| 50-64 | 353,539 (27.0%) | 37,715 (26.8%) | 19,893 (18.0%) | 41,830 (31.1%) | 32,881 (26.0%) | |
| 65-74 | 228,781 (17.5%) | 33,170 (23.5%) | 33,592 (30.5%) | 41,195 (30.6%) | 40,704 (32.2%) | |
| 75+ | 141,219 (10.8%) | 27,097 (19.2%) | 52,125 (47.3%) | 33,380 (24.8%) | 38,625 (30.6%) | |
| Sex | ||||||
| Female | 808,981 (61.7%) | 91,097 (64.7%) | 65,985 (59.8%) | 69,128 (51.3%) | 74,031 (58.6%) | |
| Male | 502,035 (38.3%) | 49,769 (35.3%) | 44,310 (40.2%) | 65,539 (48.7%) | 52,355 (41.4%) | |
| Race | ||||||
| White | 872,709 (66.6%) | 109,169 (77.5%) | 88,685 (80.4%) | 91,501 (67.9%) | 105,103 (83.2%) | |
| Asian | 121,508 (9.3%) | 7,866 (5.6%) | 4,913 (4.5%) | 13,320 (9.9%) | 6,632 (5.2%) | |
| Black or African American | 31,106 (2.4%) | 5,723 (4.1%) | 4,907 (4.4%) | 6,753 (5.0%) | 3,120 (2.5%) | |
| American Indian or Alaska Native | 8,972 (0.7%) | 1,827 (1.3%) | 1,055 (1.0%) | 1,862 (1.4%) | 881 (0.7%) | |
| Native Hawaiian or Other Pacific Islander | 5,564 (0.4%) | 875 (0.6%) | 612 (0.6%) | 1,326 (1.0%) | 453 (0.4%) | |
| Unknown | 271,157 (20.7%) | 15,406 (10.9%) | 10,123 (9.2%) | 19,905 (14.8%) | 10,197 (8.1%) | |
| Ethnicity | ||||||
| Not Hispanic or Latino | 985,681 (75.2%) | 124,093 (88.1%) | 97,030 (88.0%) | 113,182 (84.0%) | 115,697 (91.5%) | |
| Hispanic or Latino | 170,779 (13.0%) | 11,624 (8.3%) | 8,285 (7.5%) | 16,418 (12.2%) | 6,421 (5.1%) | |
| Unknown | 154,556 (11.8%) | 5,149 (3.7%) | 4,980 (4.5%) | 5,067 (3.8%) | 4,268 (3.4%) | |
Table 1: Baseline characteristics table for each of our comorbidity population along with the reference comorbidity-free general population. If a patient has multiple comorbidities, they were assigned to multiple corresponding comorbidity sub populations. Data represented are unweighted counts of patients.
Percentage of vaccinated with breakthrough case
The following Figure 1 and Table 2 summarize the association between comorbidities and the likelihood of having a COVID-19 breakthrough infection. We estimated two values: first, the percentage of the fully vaccinated population with a breakthrough infection. Second, an odds ratio which expresses how much more likely a patient with a specific comorbidity is having a breakthrough infection compared to the comorbidity-free general population. The Methods section describes details about our analysis. The results are based on weighted populations for each comorbidity which accounts for some of the differences between the people with and without the comorbidity. In addition, for visual simplicity, Figure 1 shows the percent of the population that experienced a breakthrough infection and the odds ratios of breakthrough infections for individuals with comorbidities compared to a comorbidity-free population, both calculated from weighted data (see Methods section). Table 2 includes the results for each individual population as well as the computed average comorbidity-free general population.
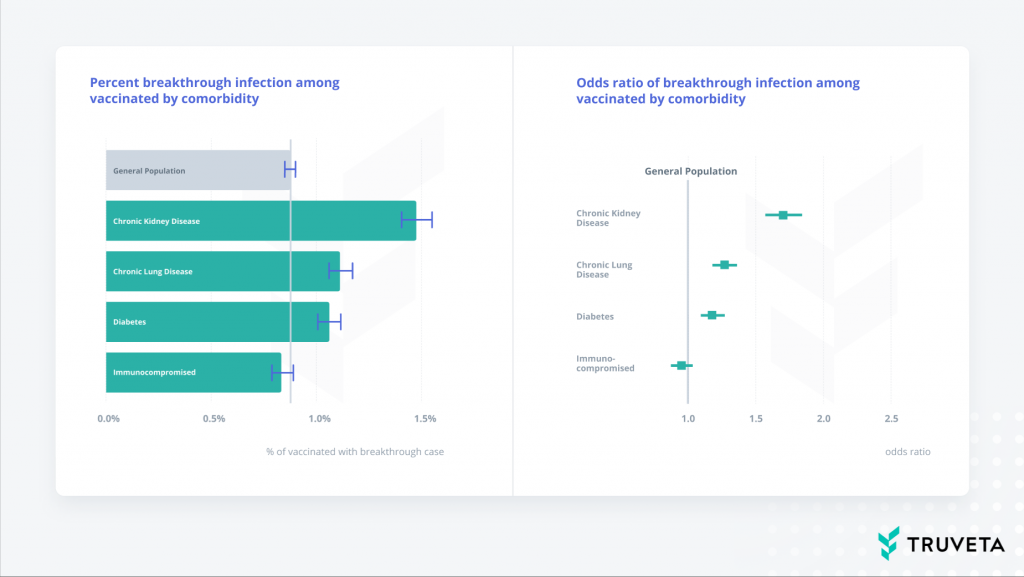
Figure 1: Left: Percentage of the analyzed vaccinated population that experienced a breakthrough COVID-19 infection among individuals with comorbidities compared to a comorbidity-free population. These values are estimated from weighted data. Right: Odds ratios for breakthrough infection associated with having a comorbidity versus having none of the analyzed comorbidities. These values are estimated from weighted data.
| Percentage of vaccinated with breakthrough case (95% CI) | |||
| Comorbidity Population | General Population | Odds Ratio | |
| Chronic Kidney Disease | 1.49% (1.42% – 1.56%) | 0.88% (0.83% – 0.94%) | 1.70 (1.57 – 1.84) |
| Chronic Lung Disease | 1.12% (1.07% – 1.18%) | 0.89% (0.84% – 0.94%) | 1.27 (1.18 – 1.37) |
| Diabetes | 1.07% (1.02% – 1.13%) | 0.91% (0.86% – 0.96%) | 1.18 (1.09 – 1.27) |
| Immunocompromised | 0.85% (0.80% – 0.90%) | 0.89% (0.84% – 0.94%) | 0.95 (0.87 – 1.03) |
| Average General Population | 0.88% (0.86% – 0.91%) | ||
Table 2: Underlying data table that is used to generate Figure 1. Each cell contains the estimated value as well as its 95% confidence interval (CI). The average comorbidity-free general population was computed to simplify the visual representation within the left side of Figure 1.
Percentage of breakthrough cases who are hospitalized
We then investigated whether comorbidities have an influence on the hospitalization rate following a breakthrough COVID-19 infection. Figure 2 and Table 3 summarize our results based on the weighted population.
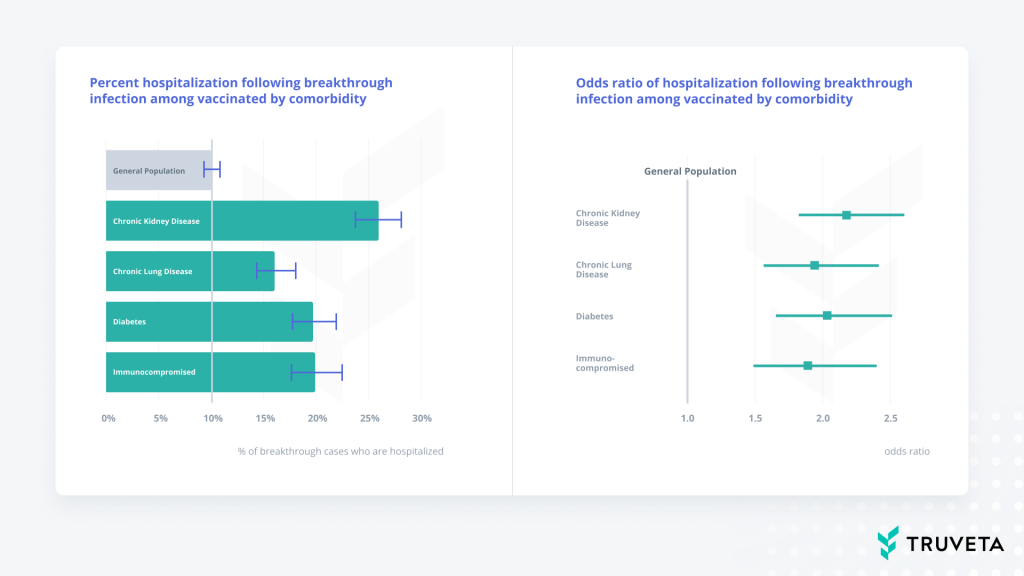
Figure 2: Left: Percentage of population that experienced hospitalization following a breakthrough COVID-19 infection among individuals with comorbidities compared to a comorbidity-free general population. These values are estimated from weighted data. Right: Odds ratios for hospitalization following a breakthrough infection associated with having a comorbidity versus having none of the analyzed comorbidities. These values are estimated from weighted data.
| Percentage of breakthrough cases who are hospitalized (95% CI) | |||
| Comorbidity Population | General Population | Odds Ratio | |
| Chronic Kidney Disease | 26.0% (23.9% – 28.2%) | 13.8% (12.2% – 15.6%) | 2.19 (1.83 – 2.62) |
| Chronic Lung Disease | 16.2% (14.5% – 18.2%) | 9.0% (7.7% – 10.6%) | 1.95 (1.57 – 2.43) |
| Diabetes | 19.8% (17.8% – 22.0%) | 10.8% (9.3% – 12.5%) | 2.04 (1.66 – 2.53) |
| Immunocompromised | 20.1% (17.7% – 22.6%) | 11.7% (9.9% – 13.8%) | 1.90 (1.50 – 2.41) |
| Average General Population | 10.2% (9.5% – 10.9%) | ||
Table 3: Each cell contains the estimated value as well as its 95% confidence interval (CI). The average comorbidity-free general population was computed to simplify the visual representation within the left side of Figure 2.
What surprised us
A few findings surprised us. It was surprising that the immunocompromised group did not demonstrate an increased risk of breakthrough infection. This is possibly due to a higher adoption of protective behaviors such as social distancing and mask wearing in this group than other groups and the general population.
This study complements other studies demonstrating increased risk for severe outcomes in these groups, although prior studies were mostly performed for unvaccinated individuals. For example, one study in an unvaccinated population reported odds ratios of greater than 2 for in-hospital mortality following COVID-19 infection in patients with diabetes, CKD, or pulmonary disease when compared with the general population (6). We identified CKD as the highest risk comorbidity. Initially we thought that this may be due to age and demographic factors, however the finding held even after adjustment for these factors. Notably, a large study in male U.S. veterans did not show an elevated risk of severe outcomes in breakthrough infections in patients with diabetes, chronic lung disease, or CKD (10). This was possibly due to their study model which matched patients by comorbidity burden thereby reducing any differences between groups. In contrast our study compared patients with identified comorbidities to a control group without these comorbidities and did not exclude or match patients with multiple comorbidities.
Why we think this is important
As vaccinated people begin to make decisions about booster vaccinations, they will be looking for information regarding their personal risk of breakthrough COVID-19 infection and severe outcomes like hospitalization. The FDA and CDC both made recommendations to include high-risk groups, such as those we studied, as groups who should receive booster vaccinations. While there is robust data showing increased risk to unvaccinated groups with comorbidities, both the FDA and CDC cited having limited data in vaccinated groups with comorbidities. The findings of this study improve the evidence and support recommendations for people with comorbidities such as chronic kidney disease, chronic lung disease, diabetes or people who are immunocompromised.
Limitations and next steps
Our study has several limitations. First, there are multiple confounding features which are difficult to address with the current data including the timing of patient vaccination, precedent COVID-19 infection, and the timing of infections in relation to SARS-CoV-2 variants of concern. In the future we hope to correct for these potential confounders. Secondly, our current data definitions rely heavily on ICD-10-CM codes, except for CKD which incorporates lab values. This limits the scope and may limit the accuracy of our comorbidity groups and COVID-19 diagnoses. For example, adding labs would improve sensitivity for COVID-19 cases. Additionally, the diagnosis-based COVID-19 definition used here may bias the underlying patient sample towards a sicker population as only individuals with sufficient symptoms will generally seek care for COVID and acquire a diagnostic code. Another limitation is that breakthrough hospitalizations were related temporally to COVID-19 infection however did not require that the hospitalization be specifically for COVID-19 infection. It was possible that individuals were hospitalized with other ailments and happened to test positive for COVID-19. This served to increase sensitivity of capturing hospitalizations following COVID-19 diagnosis however at the cost of specificity; a sensitivity analysis investigating alternative definitions for breakthrough hospitalizations, including a definition which narrows hospitalizations to those where the primary diagnosis was COVID-19, is a next step in this analysis. As more data becomes available, we plan to include SARS-CoV-2 variants of concern, geography, and more comorbidities.
There are multiple next steps to improve our statistical analysis. For example, our inverse probability weighting model currently considers the confounding effect of race, age, and sex. Future iterations should account for ethnicity, as this is an important dimension of patient demography which may be correlated with unmeasured features which could affect if a patient has a comorbidity or not.
Additionally, our current outcome models consider only the effect of a single comorbidity on our outcomes of interest (see Methods). Future iterations of this analysis should consider the effect of multiple simultaneous comorbidities on outcome as well as their interaction effects. Currently, we are treating each comorbidity as purely independent, but we know that patients can have multiple comorbidities (e.g., diabetes and CKD). In a future iteration of this analysis, we should also consider the interaction effects of patients having multiple comorbidities on probability of outcome events. For example, our model for diabetes does not consider if a patient has CKD or the other comorbidities. Future models of breakthrough infection and hospitalization outcomes should consider how the interactions among these comorbidities may contribute to differences in odds of breakthrough infection and hospitalization.
As we build our platform there will be even more we can do, with even greater rigor – but we want to start sharing and engaging now, and working with the researchers and clinicians we aim to serve to shape a shared asset that will create value for all. Please send us your feedback and ideas and help us to save lives with data.
Appendix: Methods
Our study population consisted of 1,707,650 fully vaccinated patients present on the Truveta Platform. A patient was considered fully vaccinated two weeks after receiving two mRNA vaccine doses (Moderna, Pfizer) or two weeks after receiving a single dose of the Janssen vaccine. Additionally, patients were excluded from our study population if they were missing any of sex, age, had no health system encounters following vaccination, were missing their date of being fully vaccinated, were under 12 years of age at time of vaccination, or had any vaccination events prior to December 1st, 2020. These last three criteria likely indicated a recording error in patient age at vaccination or their time of vaccination. All vaccination events were extracted from the electronic health record from member health systems; these events generally consisted of vaccinations that took place within the health system as well as vaccination records actively pulled from the health system’s respective state’s Immunization Information System.
Comorbidities were defined using Elixhauser comorbidity ICD-10-CM diagnostic codes taken from the patient’s medical record (14). Chronic kidney disease (CKD) was defined as the earliest date a patient had an ICD-10-CM diagnostic code consistent with CKD or when Kidney Disease: Improving Global Outcomes (KDIGO) criteria for CKD were met by estimated glomerular filtration rate (eGFR) criteria (15). For each comorbidity of interest, patients with that comorbidity diagnosed prior to being fully vaccinated were compared with patients with none of the comorbidities of interest. Patients who were diagnosed with a comorbidity after being fully vaccinated were excluded from analysis.
Our response variables of interest were 1) if a patient experienced a breakthrough COVID-19 infection at any point after being fully vaccinated, and 2) if a patient who experienced a breakthrough infection was hospitalized. COVID-19 infection was defined as a patient’s first diagnosis of COVID-19 as per ICD-10-CM code. Only the patient’s first COVID-19 infection was considered in this analysis. COVID-19 hospitalizations were defined as an inpatient encounter where the patient’s COVID-19 positivity fell within the interval of 14 days prior to admission up to the date of discharge. Both outcomes are binary (0/1) values.
To account for confounding demographic factors, we used inverse probability weighting for each of the comorbidities with the following confounding features as predictors: five age brackets, six race groups, and sex. Propensity scores were estimated using logistic regression and transformed into inverse probability weights to estimate the average effect of the comorbidity on those patients’ outcomes (16). Inverse probability weights were calculated separately for the breakthrough infection analysis and the hospitalization following breakthrough infection analysis.
Event rates were calculated using the weighted data. The proportion of breakthrough cases was calculated as the number of patients with and without a given comorbidity who experienced a breakthrough COVID-19 infection divided by the total number of patients with and without a given comorbidity to yield a rate, weighted by the inverse probability weights. The confidence interval for this proportion was calculated using the normal approximation of the binomial confidence interval which is suitable for large samples (17).
The weighted event rates for the population of patients without any of the comorbidities was different for each of the focal comorbidities because of how each population was weighted to resemble their respective group of patients with the comorbidity. To ease visual comparison, we calculated a composite of these four weighted event rates using a Binomial generalized linear model with only an intercept to estimate an average event rate from these samples. This composite was not used in any statistical comparisons and was only used to ease visual presentation (e.g., Figure 1). This process was repeated for patients who experienced hospitalization post breakthrough COVID-19.
Odds ratios of breakthrough associated with a comorbidity were calculated using logistic regression weighted using the calculated inverse probability weights. This method was repeated for calculating the odds ratio of hospitalization following breakthrough infection associated with a comorbidity. The confounding demographic features used in the weighting model were included as additional covariates in these regression models. Odds ratios were calculated as the exponentiated regression coefficient (18) for the comorbidity covariate in the logistic regression model. Each comorbidity was analyzed independently. This process was repeated for the analysis of hospitalization following breakthrough infection.
Analysis was done using the R programming language (4.1.1) along with the following packages: arrow, broom, dplyr, ggplot2, janitor, magrittr, purrr, questionr, rlang, stringr, tableone, targets, tibble, and tidyr (see Software References below).
References
- Puranik A, Lenehan PJ, Silvert E, Niesen MJM, Corchado-Garcia J, O’Horo JC, et al. Comparison of two highly-effective mRNA vaccines for COVID-19 during periods of Alpha and Delta variant prevalence [Internet]. Public and Global Health; 2021 Aug [cited 2021 Oct 27].
- Keehner J, Horton LE, Binkin NJ, Laurent LC, Pride D, Longhurst CA, et al. Resurgence of SARS-CoV-2 Infection in a Highly Vaccinated Health System Workforce. N Engl J Med. 2021 Sep 30;385(14):1330–2.
- Oliver, Sara. ACIP Presentation Slides: Sept 22-23, 2021, Evidence to Recommendation Framework: Pfizer-BioNTech COVID-19 Booster Dose.
- Dooling K. ACIP Presentation Slides: Oct 21, 2021, Evidence to Recommendation Framework: Moderna & Janssen COVID-19 Vaccine Booster Dose.
- Fried MW, Crawford JM, Mospan AR, Watkins SE, Munoz B, Zink RC, et al. Patient Characteristics and Outcomes of 11 721 Patients With Coronavirus Disease 2019 (COVID-19) Hospitalized Across the United States. Clin Infect Dis. 2021 May 18;72(10):e558–65.
- Mesas AE, Cavero-Redondo I, Álvarez-Bueno C, Sarriá Cabrera MA, Maffei de Andrade S, Sequí-Dominguez I, et al. Predictors of in-hospital COVID-19 mortality: A comprehensive systematic review and meta-analysis exploring differences by age, sex and health conditions. Verdonck K, editor. PLOS ONE. 2020 Nov 3;15(11):e0241742.
- Gao Y, Chen Y, Liu M, Shi S, Tian J. Impacts of immunosuppression and immunodeficiency on COVID-19: A systematic review and meta-analysis. J Infect. 2020 Aug;81(2):e93–5.
- Saini KS, Tagliamento M, Lambertini M, McNally R, Romano M, Leone M, et al. Mortality in patients with cancer and coronavirus disease 2019: A systematic review and pooled analysis of 52 studies. Eur J Cancer. 2020 Nov;139:43–50.
- Butt AA, Omer SB, Yan P, Shaikh OS, Mayr FB. SARS-CoV-2 Vaccine Effectiveness in a High-Risk National Population in a Real-World Setting. Ann Intern Med. 2021 Oct;174(10):1404–8.
- Butt AA, Yan P, Shaikh OS, Mayr FB. Outcomes among patients with breakthrough SARS-CoV-2 infection after vaccination in a high-risk national population. EClinicalMedicine. 2021 Oct;40:101117.
- Moutschen MP, Scheen AJ, Lefebvre PJ. Impaired immune responses in diabetes mellitus: analysis of the factors and mechanisms involved. Relevance to the increased susceptibility of diabetic patients to specific infections. Diabete Metab. 1992 Jun;18(3):187–201.
- O’Dwyer DN, Dickson RP, Moore BB. The Lung Microbiome, Immunity, and the Pathogenesis of Chronic Lung Disease. J Immunol. 2016 Jun 15;196(12):4839–47.
- Vaziri ND, Pahl MV, Crum A, Norris K. Effect of Uremia on Structure and Function of Immune System. J Ren Nutr. 2012 Jan;22(1):149–56.
- Elixhauser A, Steiner C, Harris DR, Coffey RM. Comorbidity Measures for Use with Administrative Data: Med Care. 1998 Jan;36(1):8–27.
- Chapter 1: Definition and classification of CKD. KDIGO 2012 Clin Pract Guidel Eval Manag Chronic Kidney Dis. 2013 Jan 1;3(1):19–62.
- Austin PC. An Introduction to Propensity Score Methods for Reducing the Effects of Confounding in Observational Studies. Multivar Behav Res. 2011 May 31;46(3):399–424.
- Agresti A. Categorical data analysis. 3rd ed. Hoboken, NJ: Wiley; 2013. 714 p. (Wiley series in probability and statistics).
- Gelman A, Hill J, Vehtari A. Regression and other stories. Cambridge, United Kingdom: Cambridge University Press; 2021. 534 p. (Analytical methods for social research).
Software References
R, R Core Team (2021). R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria.
arrow, Neal Richardson, Ian Cook, Nic Crane, Jonathan Keane, Romain François, Jeroen Ooms and Apache Arrow (2021). arrow: Integration to ‘Apache’ ‘Arrow’. R package version 5.0.0.2.
broom, David Robinson, Alex Hayes and Simon Couch (2021). broom: Convert Statistical Objects into Tidy Tibbles. R package version 0.7.9.
dplyr, Hadley Wickham, Romain François, Lionel Henry and Kirill Müller (2021). dplyr: A Grammar of Data Manipulation. R package version 1.0.7.
ggplot2, H. Wickham. ggplot2: Elegant Graphics for Data Analysis. Springer-Verlag New York, 2016.
here, Kirill Müller (2020). here: A Simpler Way to Find Your Files. R package version 1.0.1.
janitor, Sam Firke (2021). janitor: Simple Tools for Examining and Cleaning Dirty Data. R package version 2.1.0.
magrittr, Stefan Milton Bache and Hadley Wickham (2020). magrittr: A Forward-Pipe Operator for R. R package version 2.0.1.
purrr, Lionel Henry and Hadley Wickham (2020). purrr: Functional Programming Tools. R package version 0.3.4.
questionr, Julien Barnier, François Briatte and Joseph Larmarange (2021). questionr: Functions to Make Surveys Processing Easier. R package version 0.7.5.
rlang, Lionel Henry and Hadley Wickham (2021). rlang: Functions for Base Types and Core R and ‘Tidyverse’ Features. R package version 0.4.11.
stringr, Hadley Wickham (2019). stringr: Simple, Consistent Wrappers for Common String Operations. R package version 1.4.0.
tableone, Kazuki Yoshida and Alexander Bartel (2021). tableone: Create ‘Table 1’ to Describe Baseline Characteristics with or without Propensity Score Weights. R package version 0.13.0.
targets, Landau, W. M., (2021). The targets R package: a dynamic Make-like function-oriented pipeline toolkit for reproducibility and high-performance computing. Journal of Open Source Software, 6(57), 2959.
tibble, Kirill Müller and Hadley Wickham (2021). tibble: Simple Data Frames. R package version 3.1.5.
tidyr, Hadley Wickham (2021). tidyr: Tidy Messy Data. R package version 1.1.4.

